A team of researchers has developed a glowing molecule that could help the fight against antibiotic resistance, by alerting doctors to what kind of bacterial threat is affecting their patients.
Researchers at The University of Texas at Austin has developed chemical probes that can help identify an enzyme produced by some types of E. coli and pneumococcal bacteria. The enzyme is known to break down several common types of antibiotics, making these bacteria dangerously resistant to treatment.
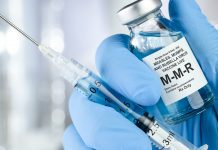
The team looked at the threat posed by the bacterial enzyme called New Delhi metallo-beta-lactamase (NDM) and set out to create a molecule that glows when it comes into contact with it. When these chemical probes are added to a test tube, they bind to the enzyme and glow – a tool that could be used to alert doctors to what kind of bacterial threat is affecting their patients and tell them which antibiotics to use.
The paper has been published in the Journal of the American Chemical Society.
A new weapon against antibiotic resistance
As well as indicating the presence of the NDM enzyme, the florescent chemical probe developed by Emily Que, assistant professor of chemistry and one of the leading researchers on the team, and Walt Fast, a professor of chemical biology and medicinal chemistry, could help in the search for alternative ways to combat these resistant bacteria. One treatment option that doctors use with resistant bacteria is to combine common antibiotics and an inhibitor – although there is no known clinically effective inhibitor for NDM-producing bacteria, Que’s probe could help find one.
Once the probe has bound to the enzyme and begun to glow, if an effective inhibitor is introduced, it will knock the probe loose and the glow would stop. This allows scientists to test a high volume of potential drugs very quickly.
Que said: “In response to antibiotic treatment, bacteria have evolved various mechanisms to resist that treatment, and one of those is to make enzymes that basically chew up the antibiotics before they can do their job. The type of tool we developed gives us critical information that could keep us one step ahead of deadly bacteria.”
“This allows us to work towards developing therapies and eventually understanding evolutionary characteristics of such proteins,” said Radhika Mehta, a recent UT Austin doctoral graduate and lead author on the paper. Mehta is currently a postdoctoral fellow in the Merchant Lab at the University of California, Berkeley.
Nutritional immunity
The study also examined the process of nutritional immunity, which comes from the human body’s production of proteins in response to an infection. The proteins take up all the available metals in the body, such as the zinc required to make NDM, making the bacteria more susceptible to attack. The probe could be used to study nutritional immunity and NDM because it will glow only in the presence of the zinc needed to form the enzyme.
“The evolution of this bacteria, since its discovery in 2008, indicates that not only is it developing antibiotic resistance, it’s attempting to combat this natural human immune process. That’s particularly scary,” Que said.